This is a practical tutorial on performing PCA on R. If you would like to understand how PCA works, please see my plain English explainer here.
Reminder: Principal Component Analysis (PCA) is a method used to reduce the number of variables in a dataset.
We are using R’s USArrests dataset, a dataset from 1973 showing, for each US state, the:
- rate per 100,000 residents of murder
- rate per 100,000 residents of rape
- rate per 100,000 residents of assault
- % of the population that is urban
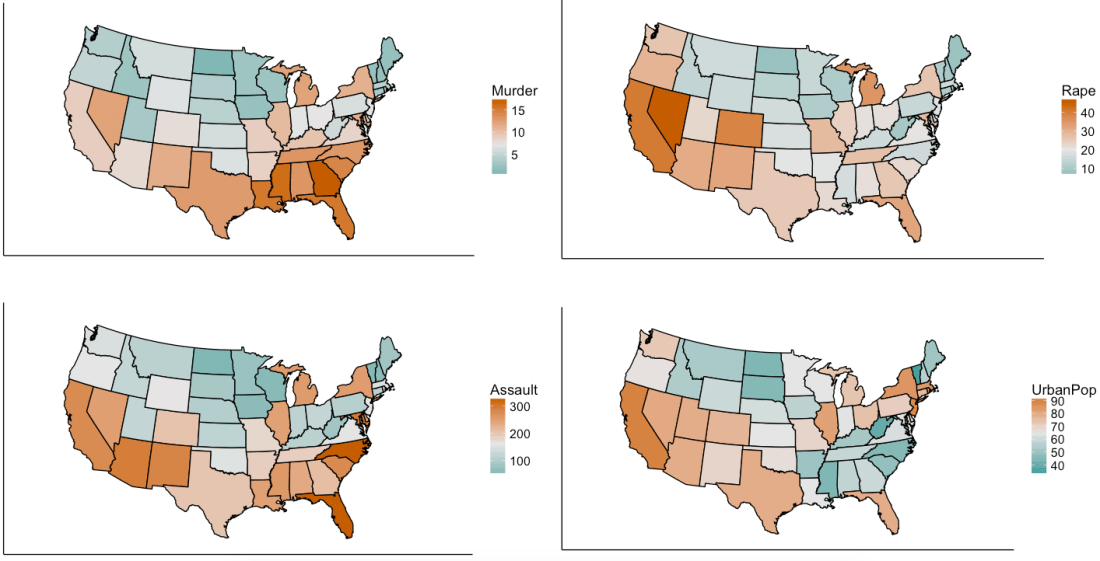
Now, we will simplify the data into two-variables data. This does not mean that we are eliminating two variables and keeping two; it means that we are replacing the four variables with two brand new ones called “principal components”.
This time we will use R’s princomp function to perform PCA.
Preamble: you will need the stats package.
Step 1: Standardize the data. You may skip this step if you would rather use princomp’s inbuilt standardization tool*.
Step 2: Run pca=princomp(USArrests, cor=TRUE) if your data needs standardizing / princomp(USArrests) if your data is already standardized.
Step 3: Now that R has computed 4 new variables (“principal components”), you can choose the two (or one, or three) principal components with the highest variances.
You can run summary(pca) to do this. The output will look like this:
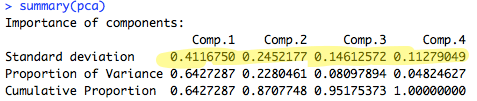
As you can see, principal components 1 and 2 have the highest standard deviation / variance, so we should use them.
Step 4: Finally, to obtain the actual principal component coordinates (“scores”) for each state, run pca$scores:

Step 5: To produce the biplot, a visualization of the principal components against the original variables, run biplot(pca):
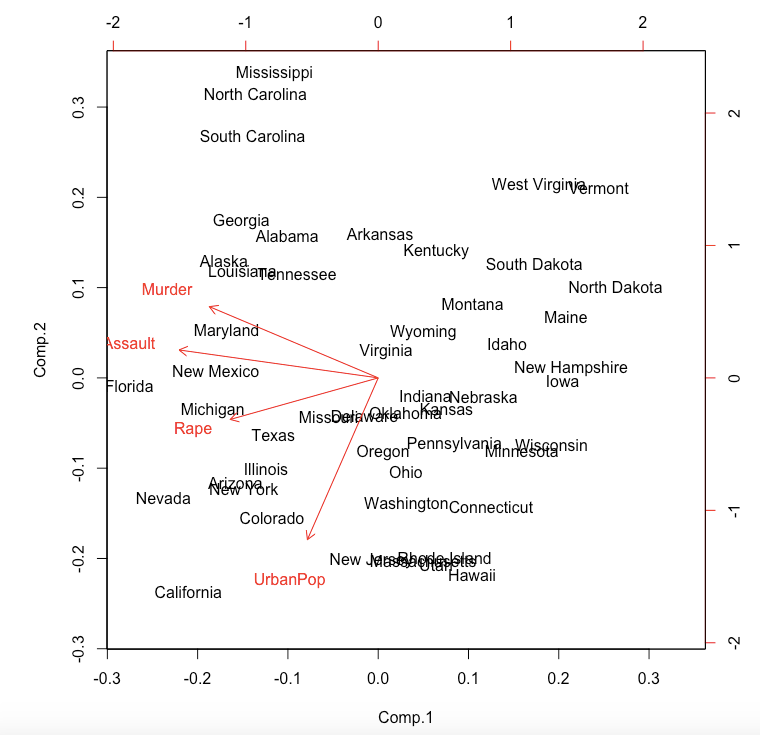
The closeness of the Murder, Assault, Rape arrows indicates that these three types of crime are, intuitively, correlated. There is also some correlation between urbanization and incidence of rape; the urbanization-murder correlation is weaker.
*princomp will turn your data into z-scores (i.e. subtract the mean, then divide by the standard deviation). But in doing so, one is not just standardizing the data, but also rescaling it. I do not see the need to rescale, so I choose to manually translate the data onto a standard range of [0,1] using the equation:

